
Novick Lab Research
Investigators in NYU Langone’s Novick Lab lead collaborative projects to advance understanding of Staphylococcus aureus and develop new methods to control, treat, or prevent infections.
In one of our projects, in collaboration with David L. Stokes, PhD, and Tom W. Muir, PhD, at Princeton University, we are attempting to determine the crystal structure of the staphylococcal agr signal receptor, AgrC, in order to define the mechanism of signal transduction by this protein. We are also working on the dynamics of agr activation in vitro at the single cell level, using fluorescent gene stags, and propose to move to an in vivo model as soon as the in vitro results are definitive.
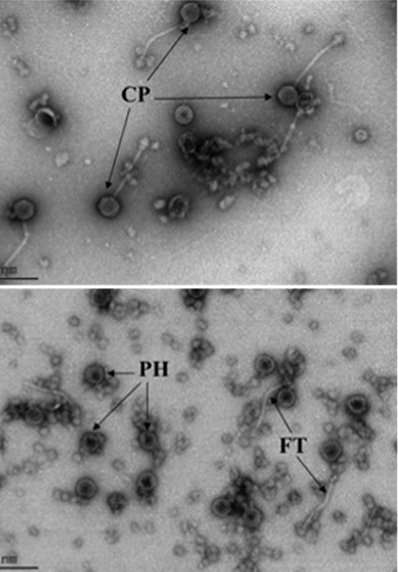
In a second project, we are studying the mechanism by which the phage-inducible mobile pathogenicity islands of S. aureus interfere with the propagation of their helper bacteriophages. These mechanisms include an island-encoded protein that inhibits the phage-encoded small terminase subunit, diversion of phage virion proteins to the formation of island-specific capsids, and inhibition of helper phage late gene transcription. The interference also results in increased survival of phage-infected cells by an unknown mechanism. A recent finding is that the mobile islands can participate in genetic transduction, a process formerly thought to be the exclusive province of bacteriophages.
A third area of concentration in our lab is the development of methods to control, treat, or prevent staphylococcal infections. These methods include analysis of agr-encoded quorum-sensing peptides as inhibitors of virulence and for their potential to modify biofilm structure. They also include the development of an antistaphylococcal vaccine and of a powerful staphylolytic enzyme. Finally, we are exploring the potential of probiotics to prevent Clostridium difficile infections owing to antibiotic depletion of the normal intestinal flora.