
Bach Lab Research
The Bach Lab is currently focused on four main areas of research, each of which uses Drosophila melanogaster as a model system.
Cell Competition
Supercompetition occurs when cells with oncogenic mutations out-compete (i.e., kill and replace) neighboring wild-type cells. Supercompetition has been compared to field cancerization, wherein a single, pre-malignant cell can clonally dominate a tissue. We showed that cells in developing Drosophila epithelia with sustained JAK/STAT signaling cause the death of neighboring wild-type cells and then increase their division rate to replace the outcompeted cells. Through RNA-seq and functional validation assays, we have identified key, differentially-expressed factors that allow STAT supercompetitors to kill wild-type neighbors. Current efforts are focused on identifying how these genes control competition.
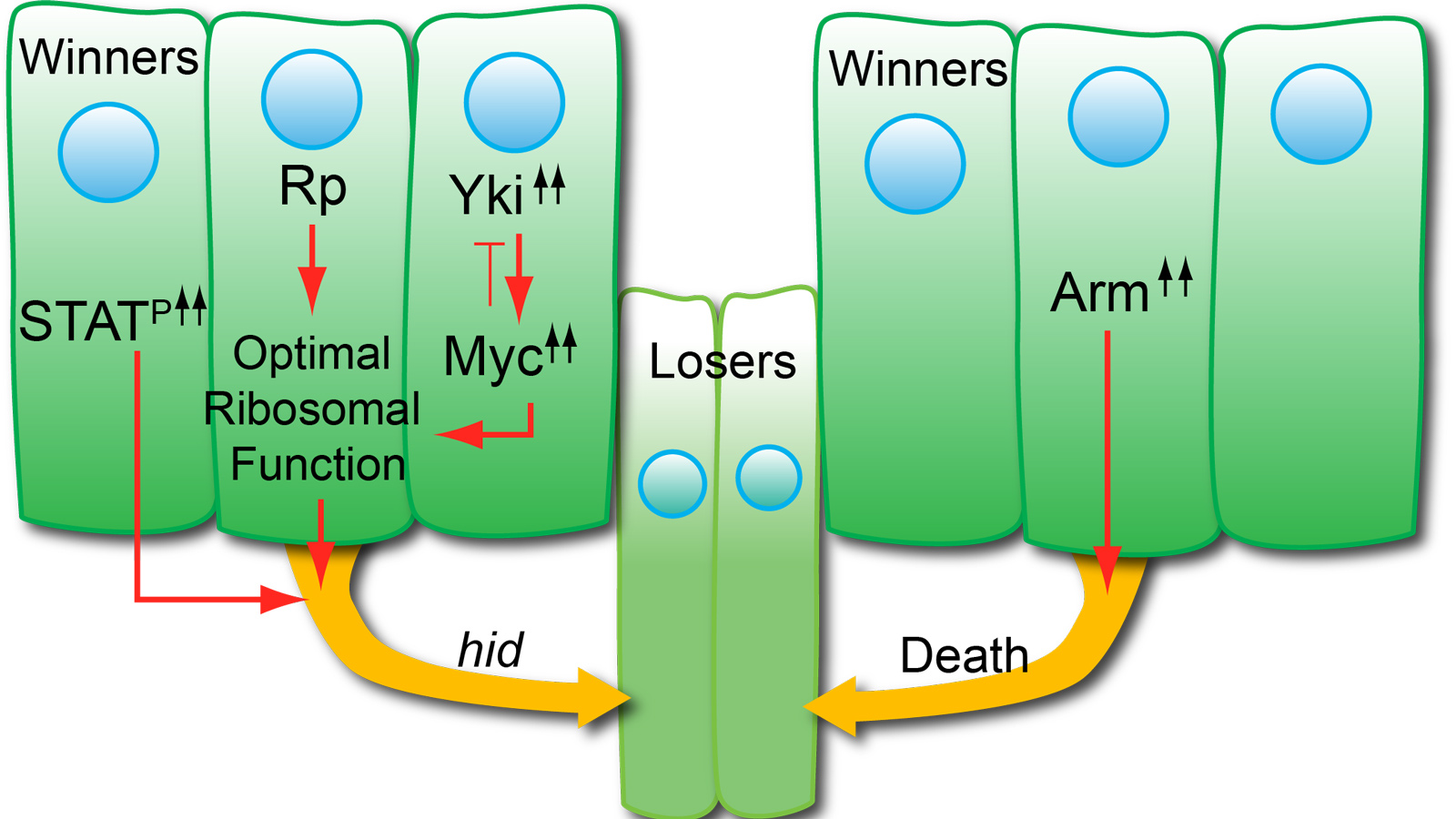
Note: In the image above, supercompetitors (orange) have acquired a mutation that increases their fitness and are able to outcompete wild-type cells (green). A clone of supercompetitors (orange) induces apoptosis (dark green) and basal extrusion (orange arrows) of surrounding wild-type cells located up to eight cell diameters away. The subsequent proliferation of supercompetitors replaces the outcompeted wild-type cells, resulting in their increased contribution to the tissue.
Stem Cell Competition
In the Drosophila testis, there is a single somatic niche that supports two populations of stem cells: germline stem cells (GSCs), which produce sperm, and somatic cyst stem cells, which produce Sertoli-like cells that support spermatogonia. We have shown that both populations compete for limited space at the niche. We have identified mutations that endow a stem cell with a competitive advantage to occupy its niche, including loss of tumor suppressors Hippo/Salvador/Warts and Patched. Through genetic approaches, we have identified additional mutations that impart a competitive advantage to stem cells, and we are keen to identify the underlying gene responsible for hyper-competition. Projects include using extended live-imaging to determine the dynamics of hyper-competitive stem cells and transcriptomics/bioinformatics to determine target genes differentially expressed in these stem cells.
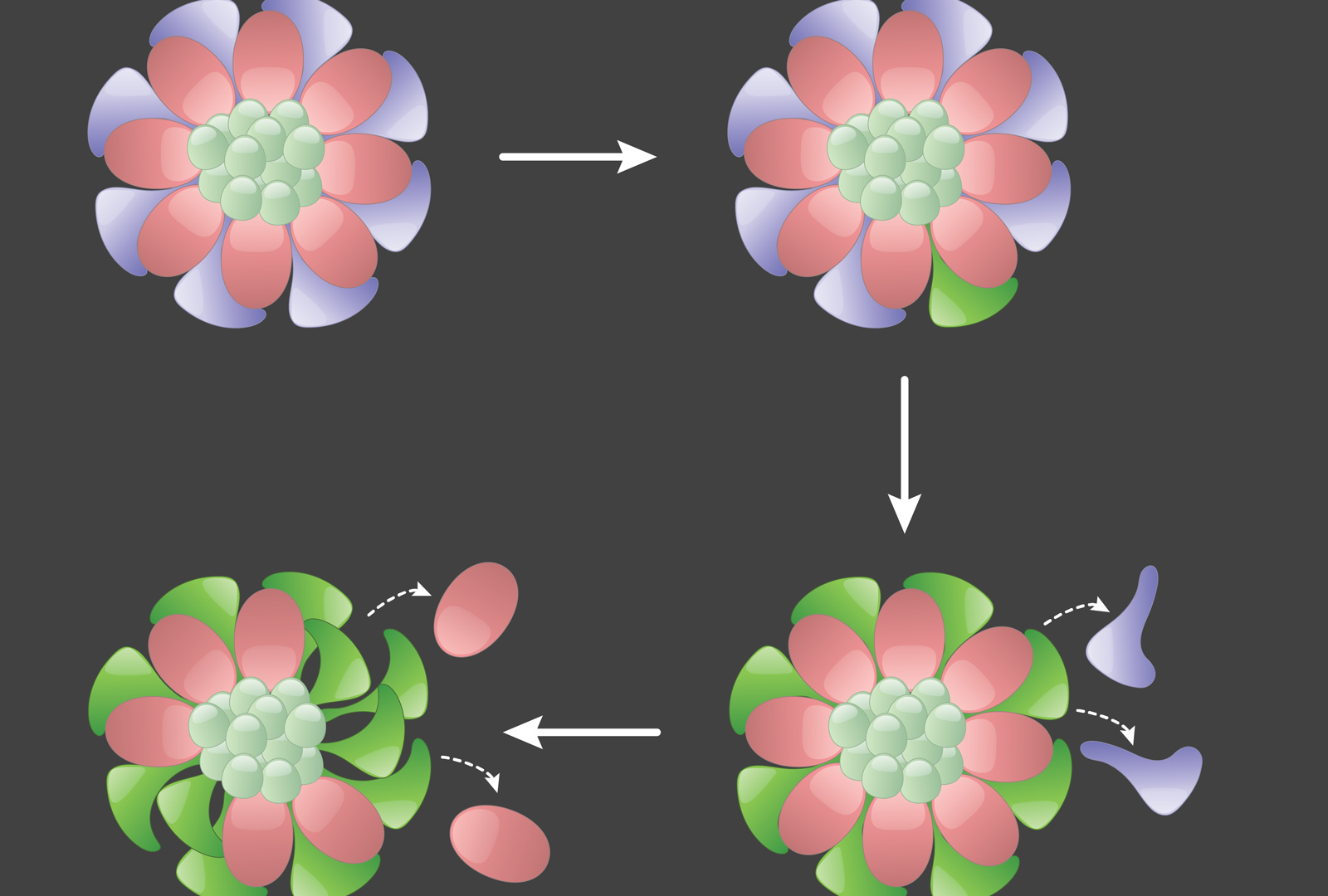
Note: In the image above, on the top left, germline stem cells (GSCs, pink) and somatic cyst stem cells (CySCs, dark blue) adhere to and are supported by the somatic niche (light green). On the top right, a labelled CySC clone (bright green) acquires a mutation that increases its fitness. On the bottom right, this mutant CySC out-competes its wild-type CySC neighbors, which leave the niche. On the bottom left, the mutant CySC also outcompetes wild-type GSCs, forcing them to leave the niche.
Dedifferentiation During Aging
We discovered that under challenging conditions, like starvation, spermatogonial dedifferentiation is critical to maintaining a robust pool of GSCs. We identified Jun N-terminal kinase (JNK) as a key signaling pathway that controls dedifferentiation and also determined that epigenetic modifications are required for this process. Additionally, the ability of spermatogonia to dedifferentiate declines steeply during aging. Current work is focused on identifying molecular targets of JNK and of epigenetic factors that are active during dedifferentiation in young and old males.
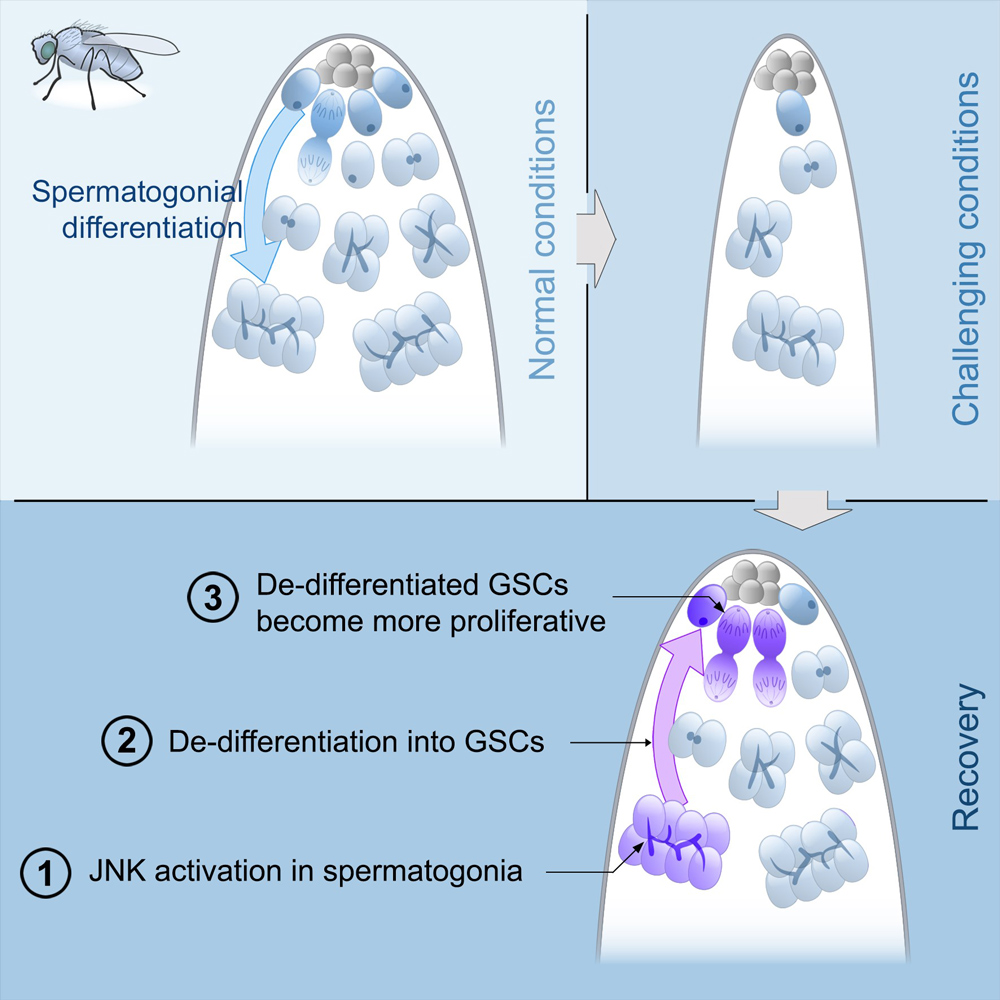
Note: In the image above, on the top left, under normal conditions, GSCs divide and their offspring undergo normal differentiation into spermatogonia. On the top right, under challenging conditions like starvation, there is a significant reduction in the number of GSCs and their offspring. On the bottom, during the recovery phase from stress, JNK signaling is triggered in spermatogonia (number 1), these cysts break apart and the germ cells dedifferentiate into GSCs (number 2). Dedifferentiated GSCs have higher proliferation rates than their wild type sibling GSCs and produce more offspring (number 3).
Sex Transformation
The production of gametes in mammals and flies requires that the sex of the germline and the sex of the soma are the same. If somatic support cells of the gonad lose their sex identity, gametogenesis collapses. We found that when somatic cells in the testis lose expression of the transcription factor Chinmo, they lose “maleness” and become feminized. This sex-transformation causes a catastrophic loss of spermatogenesis and the flies become infertile. We identified several target genes of Chinmo that repress female identity in male cells. Current efforts are aimed at determining where these target genes act in the sexual identity pathway.
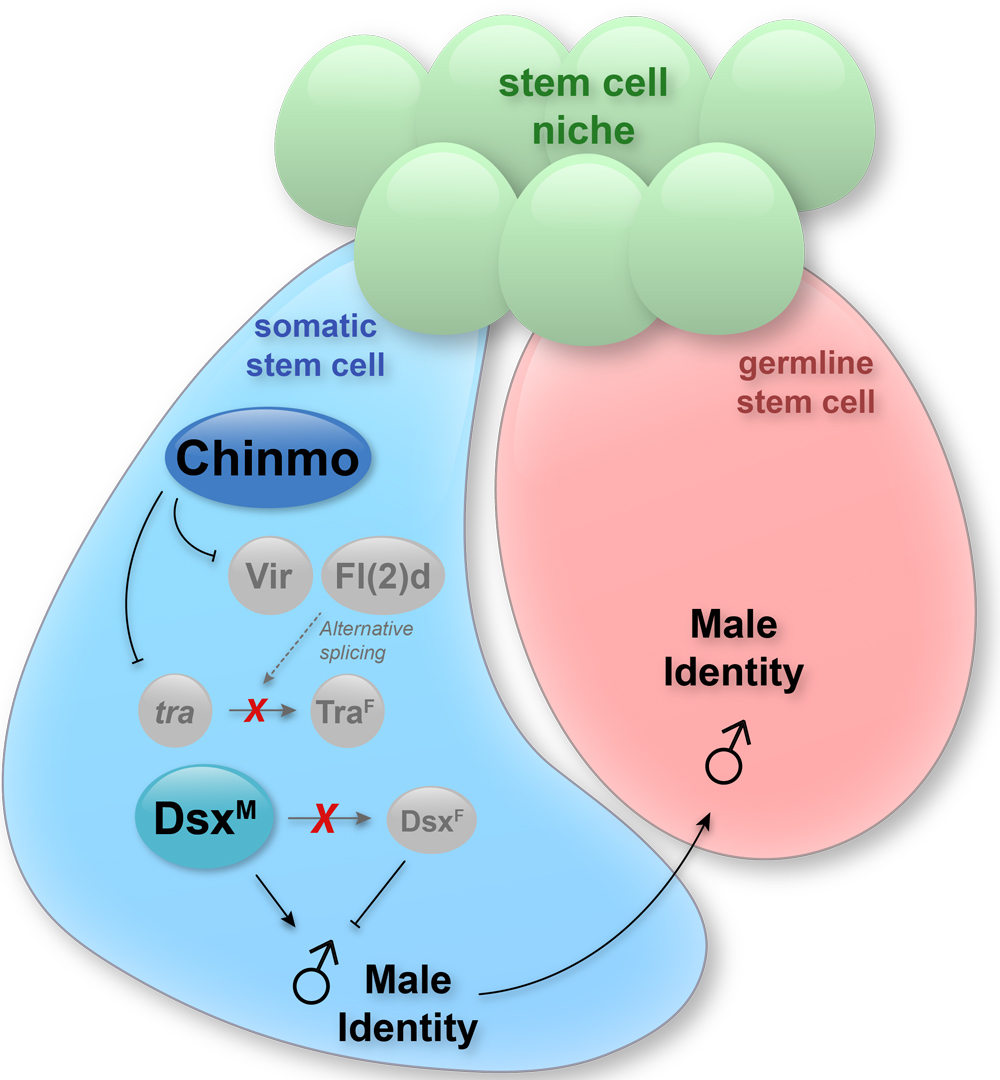
Note: In the image above, Chinmo is expressed in a sex-specific manner: it is expressed only in male but not female gonadal somatic cells. In somatic cells of the testis, Chinmo represses expression of transformer (tra), virilizer (vir) and fl(2)d. This safeguards male sexual identity by reducing the availability of tra pre-mRNA and of factors (i.e., Vir and Fl(2)d)) that can splice it into the female sex determinant traF. The presence of Chinmo in male somatic cells ensures that neither tra nor dsx pre-mRNA are alternatively spliced, resulting in the production of DsxM protein, which ensures male-specific transcription of target genes. The male sexual identity of the somatic cell must match that of germ cell for proper spermatogenesis.