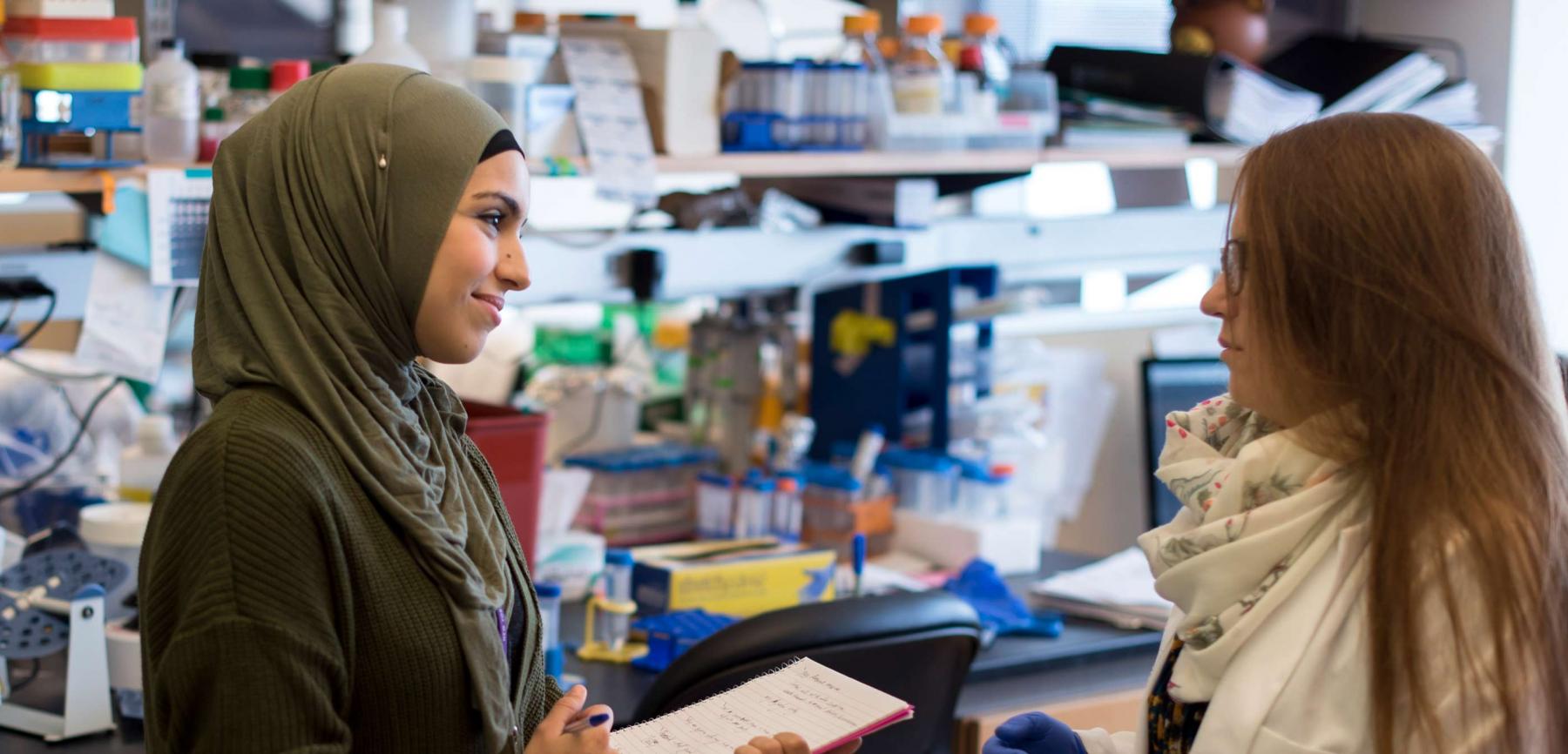

Developmental Genetics
Faculty whose research focuses on Developmental Genetics at the Vilcek Institute of Graduate Biomedical Sciences use genetic approaches to increase current understanding of developmental mechanisms.
Coursework
Explore our Faculty Mentors

Apply to Our Graduate Program

Events
Students have the opportunity to work with a variety of genetic systems including Drosophila, Caenorhabditis elegans, mouse, chicken, and zebrafish to study diverse developmental processes such as the following:
- pattern formation
- cell fate determination
- gene expression
- cell-cell signaling
- cell migration
- stem cell self-renewal
- morphogenesis
- neural circuit formation
To learn more about research into developmental genetics at NYU Grossman School of Medicine, contact one of our Pre-Candidacy Faculty Advisors or Student Representatives.
For general information about graduate programs at Vilcek Institute of Graduate Biomedical Sciences, email Vilcek-Info@NYULangone.org.
Our Research Specialties
Applying the techniques of developmental genetics to model organisms is a proven approach to revealing the pathological underpinnings of disease. Discoveries in model systems such as C. elegans, Drosophila, and zebrafish have also led to significant advances in biotechnology.
Due to shared homologies among different organisms, results obtained from genetic analysis of one model often become rapidly applicable to others. Examples include the homeobox selector genes in Drosophila and the Ras-signaling pathway in C. elegans, which have provided important insights into patterning and oncogenesis, respectively, in humans and other mammals. Further, the discovery that homeobox genes also regulate patterning in Arabidopsis adds to the increasing evidence that developmental pathways in animals and plants may overlap due to both common ancestry and convergence.
Most of the simpler organisms are also the subjects of intense genomic analysis, providing the raw material for functional genomic comparisons that are expected to reveal features of developmental processes common to all organisms.
Vertebrate Developmental Genetics
Zebrafish are easy to raise and have a short generation time of three months. Females can lay hundreds of eggs at weekly intervals, and embryos develop rapidly, with a beating heart by 24 hours. Because the developing embryos are transparent, they can be easily studied under a dissecting microscope. Rapid development of forward genetics and large-scale mutagenesis techniques has facilitated the use of zebrafish in advancing our understanding of body patterning, neurogenesis, organ development, and many other areas.
Faculty working on zebrafish include Holger Knaut, PhD, and Jesús Torres-Vázquez, PhD.
Mice are the organisms of choice for a mammalian studies. Their relatively short generation time, small size, and large litters offer researchers several important advantages over other mammalian model organisms. The availability of inbred strains and amenability to genetic manipulation, such as gene targeting and production of transgenics, provide additional advantages. Mice have been used to further our knowledge in many areas, including brain development and human disease.
Faculty working on mice include Iannis Aifantis, PhD; Steven Burden, PhD; Jeremy Dasen, PhD; Esteban O. Mazzoni, PhD; Markus Schober, PhD; and Susan Schwab, PhD.
Invertebrate Developmental Genetics
C. elegans is a free-living, nonpathogenic nematode that normally inhabits the soil. Although primitive, this organism exhibits fundamental biological processes common to humans, thereby providing researchers with an invaluable model of study. The cell lineage of C. elegans is largely invariant, and the transparency of the worm reveals each nucleus in the body when viewed under a microscope at high power, allowing the entire lineage to be traced. Genetic manipulations, both forward and reverse, are well developed. Of note, C. elegans was the first multicellular organism to have a fully sequenced genome.
Faculty working on worms include E. Jane Hubbard, PhD and Niels Ringstad, PhD.
Drosophila is commonly known as the fruit fly. It has been used as a model system in biology since 1909. Its tremendous history provides an extensive database of genes and mutants that has now been extended by the complete genome sequence. Powerful forward genetics coupled with the ability to make transgenic flies through transposable elements provide a model system that has facilitated our understanding of everything from body patterning to organ development to behavior.
Faculty working on flies include Erika Bach, PhD; Claude Desplan, PhD; Christine Rushlow, PhD; Hyung Don Ryoo, PhD; and Jessica Treisman, PhD.
Ciona intestinalis is an ascidian, a marine invertebrate chordate that belongs to the Tunicate phylum, the closest living relative to the vertebrates. The genome has been sequenced, facilitating molecular studies.
Faculty working on Ciona Anna Di Gregorio, PhD.
The yeast Saccharomyces cerevisiae is a single-cell fungal micro-organism and was the first yeast genome to be sequenced. This organism is key in baking, wine, and beer fermentation. While yeasts in the wild are diploid or polyploid, laboratory strains have mutations that keep them as stable haploids. This allows for easy genetic analysis and manipulation, and for mating and induction of meiosis. Because S. cerevisiae is easily amenable to genetic manipulation, it is an ideal model organism for study.
Faculty working on yeast include Karim-Jean Armache, PhD.
B. subtilis is a nonpathogenic gram-positive bacterium. It is usually found in the soil, where it can promote plant growth. The biology of B. subtilis spans many processes of interest, including biofilm formation and sporulation. Sporulation is a developmental program triggered by nutrient deprivation, which leads to the formation of metabolically dormant but highly resistant cells called spores. B. subtilis cells are naturally competent, which facilitates the genetic manipulation of the bacterium.
Faculty working on B. subtilis include Patrick Eichenberger, PhD.
